- Method
- Open access
- Published:
DENA: training an authentic neural network model using Nanopore sequencing data of Arabidopsis transcripts for detection and quantification of N6-methyladenosine on RNA
Genome Biology volume 23, Article number: 25 (2022)
Abstract
Models developed using Nanopore direct RNA sequencing data from in vitro synthetic RNA with all adenosine replaced by N6-methyladenosine (m6A) are likely distorted due to superimposed signals from saturated m6A residues. Here, we develop a neural network, DENA, for m6A quantification using the sequencing data of in vivo transcripts from Arabidopsis. DENA identifies 90% of miCLIP-detected m6A sites in Arabidopsis and obtains modification rates in human consistent to those found by SCARLET, demonstrating its robustness across species. We sequence the transcriptome of two additional m6A-deficient Arabidopsis, mtb and fip37-4, using Nanopore and evaluate their single-nucleotide m6A profiles using DENA.
Background
N6-methyladenosine (m6A) is the most abundant modification found in messenger RNA (mRNA) [1, 2]. Previous studies demonstrated that m6A affected RNA processes, such as pre-mRNA splicing [3, 4], RNA stability [5], and RNA localization [6]. Recently, Nanopore direct RNA sequencing (direct RNA-Seq) makes it possible to directly “visualize” signals of RNA m6A modification. Direct RNA-Seq studies on in vitro synthetic RNAs with adenosine replaced with N6-methyladenosine triphosphate substrates observed signaling shifts between regular “A” residues and m6A residues [7] and increased base-calling errors around m6A residues [8]. Similarly, previous study also observed signaling shifts and base-calling errors between in vivo transcribed RNAs from wild-type and ime4∆ yeast mutant that lacked the m6A methyltransferase activity for RNA methylation [8]. Progress was made in locating m6A modification on RNAs using based-calling errors found in direct RNA-Seq reads from wild-type in comparison to those from m6A-deficient mutants. Taking advantage of this approach, several tools, such as differr [9], DRUMMER [10], ELIGOS [11], and xPore [12], were developed for m6A detection. However, this approach is hindered by the requirement of comparison of wild-type and m6A-deficient mutants, and by the low sensitivity for hypomethylated m6A sites.
A more robust approach is to recognize m6A residues based on their unique electrical signal fingerprints in direct RNA-Seq data, which does not require m6A-deficient mutant and can pinpoint m6A residues on a given RNA molecule. For example, MINES [13] established a random forest model from ionic electrical value to call m6A events in sequence context of “AGACT”, “GGACA”, “GGACC”, and “GGACT”. Using direct RNA-Seq data generated from in vitro synthetic transcripts, several machine learning models were developed, such as EpiNano that uses support vector machines, and Nanom6A that uses Extreme Gradient Boosting [14]. Both can predict RNA m6A sites without the requirement of m6A-deficient mutants. However, some drawbacks in these methods hinder their performance on m6A prediction. For one, these methods were trained with the direct RNA-Seq data from in vitro synthetic transcripts, in which all “A” residues are replaced by m6A. Often the occurrences of clustered multiple m6A residues, especially consecutive m6A residues, may cause superimposed effect on signal readout. Thus, these models may not conform to those for studying m6A modification in in vivo transcribed RNA in live cells and are limited in performance on natural samples. In addition, the training data generated from in vitro synthetic transcripts had limited numbers of variants in sequence context. For example, Nanom6A were trained with direct RNA-Seq data that contains 130 sites of “RRACH” motif that is substrate for N6-methyltransferase complex. They may not be sufficient to train the deep-learning models for m6A detection, as previous studies demonstrated that the performance on prediction can be improved on sophisticated deep neural networks [15, 16] using extended training data covering more diverse combinations of the five-residue motifs.
Here, we designed a novel neural network called DENA (Deeplearning Explore Nanopore m6A) by training on direct RNA-Seq data of in vivo transcribed mRNAs from wild-type and m6A-deficient Arabidopsis thaliana (A. thaliana). Compared to the in vitro synthetic RNA data, the in vivo transcribed RNA data are more productive in building reliable models on detecting m6A, as they (1) do not contain clustered multiple m6A residues that may distort the m6A prediction model; and (2) extend coverage on more diverse combinations of the “RRACH” motifs in direct RNA-Seq data. DENA is shown to achieve accurate identification and quantification of m6A at single-nucleotide resolution in both A. thaliana and Homo sapiens. Importantly, DENA is able to detect m6A events on different isoforms of single genes (at a single-isoform level), which is unavailable in previous methods. Furthermore, we evaluated DENA in two m6A-deficient A. thaliana mutants, fip37-4 and mtb, and generated a rich resource for profiling m6A modification at single-nucleotide resolution for these A. thaliana lines. Our study established an approach to train with data containing naturally occurring m6A patterns from direct RNA-Seq sequencing of in vivo transcribed transcripts and will provide a framework for identifying other types of RNA modifications using Nanopore direct RNA sequencing.
Results and discussion
Modeling with in in vitro transcripts of saturated m6A does not conform to that of m6A modifications in in vivo transcripts
Several models [8, 14] have been developed for RNA m6A identification utilizing the training data of in vitro transcripts. Nevertheless, there is a key limitation to consider. In the twelve possible five-mers of “RRACH” motif, ten of them contain at least two adenosine residues. However, all “A” residues are replaced with m6A residues in in vitro synthetic training data, causing the aggregation of multiple m6A residues, which is likely not to occur in in vivo transcripts from live cells. Thus, we investigate the difference between the cluster m6A residues in in vitro transcripts and the naturally occurring m6A residues in in vivo transcripts on electrical signals and base call accuracy around modification sites.
In in vitro synthetic transcripts, the shift of electrical signals [7] and the enrichment of the mismatches [8] around m6A residues have been observed in modified direct RNA-Seq reads relative to unmodified reads. We aligned the in vitro synthetic reads and in vivo transcribed reads to the corresponding reference, respectively (Fig. 1a, d). We found the existence of clustered multiple mismatches within or around “RRACH” five-mers in m6A-modified reads of synthetic data (Fig. 1a). For example, in two “AAACC” five-mers from in vitro synthetic sequences, we observed distinguishing signals and consecutive mismatches within the “AAACC” pattern and its surrounding “A” residues (Fig. 1b). However, this situation was infrequent in direct RNA-Seq data of in vivo transcribed RNAs from A. thaliana (Fig. 1d), and only sporadic distinction of signals and mismatches were observed around the m6A sites. For example, only single position occurs significant mismatches around two m6A sites, which are identified by differr [9] and contained in the m6A peak from MeRIP-Seq [17, 18] (Fig. 1c, e). That is to say, the saturated m6A modification in in vitro synthetic transcripts led to different patterns of electrical signals and mismatches, relative to the m6A modification on in vivo transcripts. In addition, previous studies demonstrated that sophisticated neural networks can improve the performance on the prediction of chemical modifications [15, 16]. As mentioned in “Introduction,” the direct RNA-Seq data from in vitro synthetic transcripts had a limited number of variants in sequence context, maybe insufficient for training deep-learning models for m6A detection.
The distinction of electrical signals and mismatches between in vitro and in vivo transcripts. a The alignments are performed between the in vitro synthetic m6A-modified and unmodified reads and transcriptome reference. b The distributions of electrical signals and base-calling “errors” in in vitro reads at two “AAACC” sites. c The m6A peaks on AT2G20690 identified by two MeRIP-Seq datasets. d The alignments are generated by mapping the reads of VIRc and vir-1 to Araport11 reference in the region of AT2G20690.1. e The distributions of electrical signals and base-calling “errors” at two m6A sites identified by differr tool in native direct RNA-Seq reads of VIRc and vir-1
Identification of m6A sites in A. thaliana for subsequent neural network training
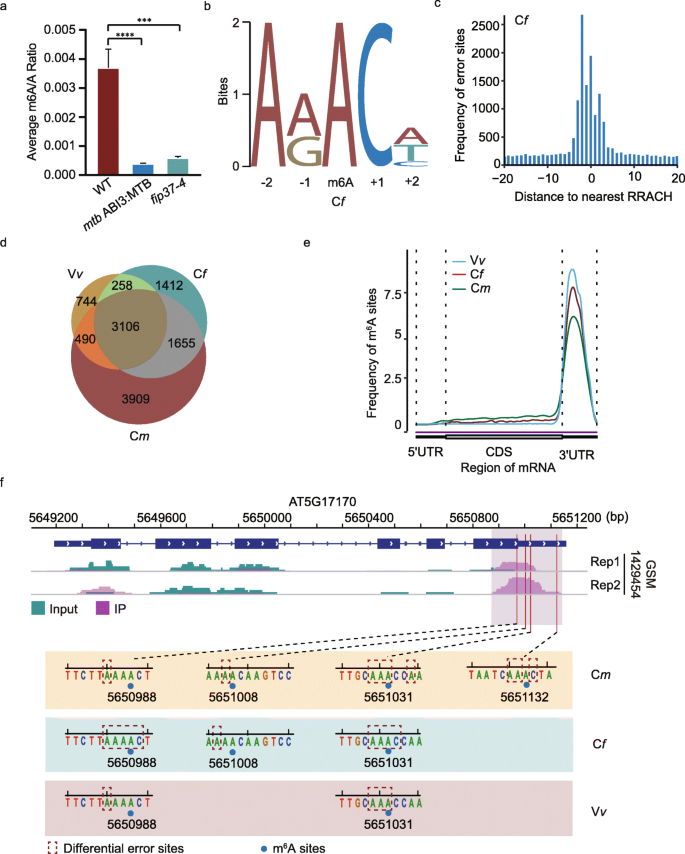
In A. thaliana, m6A modification formed with recruitment of the m6A “writer” complex containing MTA (orthologue of human methyltransferase-like3, METTL3) [19], FIP37 (orthologue of human Wilms tumor 1-associated protein, WTAP) [17, 20], MTB (orthologue of human methyltransferase-like14, METTL14) [20], and VIRILIZER [20]. We confirmed the significant decrease of m6A level in total RNA from two m6A-deficient mutants, mtb and fip37-4, using LC-MS/MS (“Methods”), and performed direct RNA-Seq on the mRNAs extracted from Col-0, mtb, and fip37-4, respectively (Fig. 2a). For each sample, over 1.8 million base-calling reads were aligned with the genome reference (TAIR10) of A. thaliana successfully (Additional file 1: Table S1), and their read-length were distributed within 800–1000 nucleotides (Additional file 1: Fig. S1a). In addition, the T-DNA insertions within FIP37 (AT3G54170) and MTB (AT4G09980) were also confirmed by direct RNA-Seq reads, respectively (Additional file 1: Fig. S1b).
Quantification of m6A levels using LC-MS/MS and identification of m6A sites using differr from direct RNA-Seq reads. a LC-MS/MS quantify the ratio of “m6A” and “A” residues from total RNA of Col-0, fip37-4, and mtb, respectively. b The motif is enriched in Cf group. c The distribution of distances between differential sites and its nearest “RRACH” motif in Cf group. d Venn diagram shows the common m6A sites identified among Cm, Cf, and Vv groups. e The m6A distribution of three groups on transcripts, are strong enriched around the stop codon. f The m6A peaks detected by MeRIP-seq and the m6A sites identified by differr in ENH1 (AT5G17170) gene from three groups, respectively
To obtain reliable m6A sites for subsequent neural network training, we used differr tool to compare the “mismatch” events in Col-0 with those in mtb (designated Cm) and with those in fip37 (designated Cf) mutants, respectively. Searching for the motifs surrounding the differential sites detected by differr, we found the most frequently identified motifs in both Cf and Cm sets closely resembled the “RRACH” (R = A/G, H = A/C/U) motif established by MeRIP-Seq in A. thaliana [17, 18, 21] (Fig. 2b; Additional file 1: Fig. S1c). These differential sites mainly fell within five nucleotides of the nearest “RRACH” motif closest to them (Fig. 2c; Additional file 1: Fig. S1d). In all, 6431 and 9160 m6A sites were detected in Cf and Cm using differr, respectively.
We further analyzed the direct RNA-Seq data from VIR-complemented (VIRc) and vir-1 mutant (vir-1) [9] (designated Vv), using the same procedure (Additional file 1: Fig. S1e, f). A total of 3106 m6A sites were common among Cm, Cf, and Vv sets, accounting about 67.6% of m6A sites in the Vv group (Fig. 2d). A strong enrichment of m6A sites around the stop codon was observed in all three groups (Fig. 2e). For example, differr detected four m6A sites in ENH1 (AT5G17170) and all of them fell within the established peaks from MeRIP-seq [18] (Fig. 2f). Altogether, these results indicated these m6A sites identified by differr were reliable. The 3106 overlapped m6A sites were used for subsequent neural network training.
Training a novel model with neural network for RNA m6A detection
The “error” events can occur at the adjacent bases of m6A sites due to the characteristic current blockade caused by m6A residues on RNA when direct RNA-Seq is performed. We asked whether we were able to separate the in vivo direct RNA-Seq reads of A. thaliana basing on the error events that occurred within the nucleotide of m6A site and five nucleotides of upstream and downstream (Fig. 3a). We extracted native m6A-modified and unmodified data of the 3106 m6A sites (shared by Cf, Cm, and Vv) from VIRc and vir- 1[9], respectively (Figs. 1d and 3a). In all, we obtained the 2,251,835 matrices of features (mean, median, standard deviation, dwell time, and base quality) from the events of direct RNA-Seq reads, which contains 507,066 in m6A-set and 1,744,769 in A-set (“Methods”). We divided them into training and testing datasets at a ratio of 7:3 for training a neural network model, named DENA (Fig. 3a). We evaluated the performance of our model, and for 12 possible five-mers of “RRACH” motif, the area under the curve (AUC) ranges from 0.90 to 0.97, and the accuracy from 0.83 to 0.93 in the testing dataset, indicating consistent performance among the twelve motifs (Fig. 3b; Additional file 1: Fig. S2a; Additional file 1: Table S2). We applied DENA to the unmodified direct RNA-Seq data of the in vitro synthetic transcripts to evaluate its false positive. The background noise was found to be predominantly below 0.1, and almost all below 0.2 (Additional file 1: Fig. S2b). Thus, we designated the m6A sites as those above the baseline value 0.1, and the high-confidence m6A sites as those above 0.2 (see “Methods” section).
Extracted native training data and trained a neural network model. a Flowcharts illustrating the classification of modified and unmodified native reads and the procedures of training DENA. b The ROC curves of DENA in the 12 consensus sequences from “RRACH” motif. c Bar plots demonstrate the reduction of m6A rates identified by DENA at the ACTB-1217 site from direct RNA-Seq data of wild-type and METTL3-knockout human cells, respectively. d Venn diagram of the number of m6A sites identified by DENA from these sites that are supported at least 50 reads in all replicates of wild-type A. thaliana, respectively. e The cross-correlation coefficient of m6A rates identified by DENA among four replicates of wild-type A. thaliana
To evaluate the ability of DENA in m6A detection and quantification, we performed predictions at the ACTB-1217 (NM_001101-1216) site, which was a known m6A site identified by the SCARLET [22] method previously, in direct RNA-Seq data of wild-type (WT) and METTL3 knockout (M3KO) human cells [23]. The modification rate at this site was 0.180 and 0.044 in WT and M3KO, respectively, suggesting a significant reduction of m6A modification in METTL3 knockout (Fig. 3c). The DENA-identified rate in WT (0.180) was close to 0.21 that was identified by SCARLET, confirming the reliability of DENA.
To check whether DENA can reproduce m6A sites among replicated experiments, we tested DENA on the four replicates of wild-type A. thaliana [9]. To reduce the impact of the sequencing depth, we considered 22729 “RRACH” sites that were supported by at least 50 direct RNA-Seq reads in each of four replicates. A total of 6591, 7484, 6722, and 6681 m6A sites were obtained in the four replicates, respectively, and 5082 sites were common among them (Fig. 3d). We computed the cross-correlation coefficient of the modification rate at these sites across four replicates, and obtained a correlation with 0.96 (Fig. 3e). Altogether, these results confirmed the reliability of DENA for m6A detection and quantification.
Validating the robustness of DENA in m6A detection in A. thaliana and human
To validate the ability of DENA in mapping m6A sites to transcriptome with single-nucleotide precision, we applied it to three published A. thaliana samples [9], including Col-0 (wild-type), VIRc, and vir-1, respectively. We scanned all “RRACH” sites on the transcriptome in three samples. 149136, 173730, and 166300 positions were supported by at least 50 direct RNA-Seq reads in Col-0, VIRc, and vir-1, respectively. A total of 46,500 (31.18%), 48,602 (27.98%), and 39,241 (23.60%) m6A sites were identified by DENA in Col-0, VIRc, and vir-1, respectively, and 25,283 sites were overlapped across all samples (Fig. 4a). A significant reduction of m6A modification rate was observed in vir-1 compared to those in both Col-0 and VIRc (Fig. 4b; Additional file 1: Fig. S3a). Importantly, the high correlation of modification rate was found between VIRc and Col-0, confirming the reliability of the m6A quantification using DENA (Fig. 4c). The m6A sites identified by DENA enriched near the stop codon and 3′UTR in all samples (Additional file 1: Fig. S3b), agreeing with the distribution of m6A modification on mRNAs.
The m6A identification for A. thaliana and human using DENA. a Venn diagram shows the common m6A sites identified by DENA among Col-0, VIRc and vir-1 samples. b Jointplot shows the correlation of m6A rates at 28299 overlapped sites between Col-0 and vir-1. c Jointplot shows the correlation of m6A modification rate at 35,048 overlapped sites between Col-0 and VIRc. d Venn diagram shows the common m6A sites identified by DENA from wild-type (WT) and METTL3-knockout (M3KO) human cells. e The m6A distribution on transcripts in WT and M3KO, respectively. f Jointplot shows the correlation of m6A rates at 7036 overlapped sites between WT and M3KO. g Pie charts display the performance of DENA on 4201 miCLIP-identified m6A sites that are supported at least 50 reads, and 3768 sites out of them are regarded as m6A modification sites by DENA. h Flowcharts illustrating the identification of m6A sites using Bst 2.0 DNA polymerase Splint ligase with qPCR. Dp and Up are down probe and up probe, respectively. i The real-time fluorescence curves produced at Site_7340 and Site_7207 on the mRNA of PRP8A, respectively. A is Site_7207 and m6A is Site_7340 on PRP8A. j Bar plot demonstrates consistency of m6A rate at the four known modified sites between DENA detection and SCARLET identification, and the significant overestimation in the prediction from Nanom6A
We further assessed the DENA robustness on other species by applying it to human direct RNA-Seq data [23]. We obtained 12,860 and 10,611 m6A sites in WT and METTL3 knockout (M3KO) samples, respectively, and 7036 sites were common between them (Fig. 4d). These m6A sites concentrated around the stop codon and 3′UTRs (Fig. 4e). Notably, the modification rate in M3KO were significantly reduced relative to that in WT (Fig. 4f).
We next compared DENA with Nanom6 A[14] that was recently developed using the in vitro synthetic training data, using the 4685 miCLIP sites identified in previous work [9]. Among the 4685 m6A sites, 4201 had a sequencing depth of at least 50, of which about 90% (3768/4201) were detected as m6A modification by DENA (Fig. 4 g), a significant improvement compared with the 66% reported by Nanom6A. Meanwhile, we further randomly selected five DENA-predicted m6A sites (PRP8A_Site_7340, CURT1B_Site_1121, EMB1467_Site_2629, NACA3_Site_802 and RPL17B_Site_688) on mRNAs of Arabidopsis identified by the SELECT experimental method [24], which utilizes the fact that m6A hinders both the single-base elongation activity of Bst DNA polymerase and the nick ligation efficiency of SplintR ligase (Fig. 4 h). For example, DENA identified a m6A site at 7340th (Site_7340) nucleotide on the PRP8A transcript (AT1G80070.1). Using the nearby unmodified “A” (Site_7207) as a control to keep the same amount of RNA template, we performed an identification on this candidate m6A-modified site (Site_7340) with SELECT method. After performing simple qPCR-based quantification of the ligation products at Site_7340 site versus nearby Site_7207 site, we found that the Ct (threshold cycle) value at Site_7340 site was significantly increased relative to that at Site_7207, indicating that the amount of final ligation products formed at Site_7340 site was dramatically reduced compared to products formed at Site_7207, confirming the m6A modification at Site_7340 (Fig. 4i). For the other four selected sites, we also detected the significant increase of Ct value at candidate m6A-modified site compared to that at nearby control “A” site, confirming the presence of m6A modification at these sites (Additional file 1: Fig. S3c). Therefore, these results confirmed the reliable prediction of m6A modification using DENA in Arabidopsis.
For human data, the m6A modification rate of several sites on two long noncoding RNAs (lncRNAs) (MALAT1 and TUG1) and three mRNAs (ACTB, TPT1, and BSG) were quantified by SCARLET method [22]. However, we found the Nanopore sequencing coverage of two lncRNAs were very low, both are fewer than 5 in all the datasets, which made it impossible to conduct a reliable analysis. We speculate about its cause and believe it is likely lncRNAs were not retained during the purification of mRNA using poly(T) before nanopore direct RNA sequencing, as the poly(A) structure may not be there for lncRNAs [25]. We thus compared DENA with Nanom6A at the known sites identified by SCARLET method on TPT1 (NM_003295), ACTB (NM_001101), and BSG (NM_198591) mRNAs [22]. The m6A rates quantified by DENA were more consistent with those by SCARLET, compared with those estimated by Nanom6A (Additional file 1: Table S3). For example, the NM_001101-1216 and NM_003295-687 sites were detected as m6A sites by DENA with 0.18 and 0.10 modification rates, respectively, which were close to the 0.21 and 0.15 quantified by SCARLET. However, m6A rates of 0.64 and 0.42 were obtained by Nanom6A at NM_001101-1216 and NM_003295-687, respectively (Fig. 4j). For NM_003295-694 and NM_198591-1442 sites, DNEA obtained 0.04 and 0.04 modification rates, respectively, which were consistent with the value of 0.04 and 0.01 by SCARLET. However, Nanom6A returned 0.12 and 0.18 modification rates at NM_003295-694 and NM_198591-1442, respectively. These results suggested the significant overestimation of modification rate by Nanom6A relative to those by SCARLET (Fig. 4j). We also investigated the m6A rate at these sites in M3KO cells using DENA. Notably, we observed significant reduction of modification rates at NM_001101-1216 and NM_003295-687 sites and slight decrease at NM_003295-694 and NM_198591-1442 compared to those in wild-type (Additional file 1: Fig. S3d). In addition, we further compared the predicted modification rates from DENA with those from other methods containing xPore, LEAD-m6A-seq and Deoxyribozyme-based Method, at the identified m6A sites in previous work [12, 26, 27] (Additional file 1: Table S3). Importantly, we can observe that the modification rates predicted by DENA are consistent with those identified by experimental methods at most sites. Concretely, the modification rates predicted by DENA at seven sites, ACTB_5527743, BSG_583239, BSG_583346, TPT1_45337310, TPT1_45337303, YTHDF2_28743593, and PARP1_226361173, are in general agreement with those verified by at least one experimental method, SCARLET and/or LEAD-m6A-seq. For example, NM_001618.4-3496, a negative site on PARP1 gene used to confirm the ability of LEAD-m6A-seq in previous work [26], was identified as non-m6A site with only 2% modification rate by LEAD-m6A-seq, which was comparable to the 7% modification rate predicted by DENA, while 44% modification rate was predicted by Nanom6A. In addition, the prediction of reduction in m6A modification rates in METTL3 knock-down mutant line using the three models, Nanom6A, xPore, and DENA, were in general agreement (Additional file 1: Table S3).
Collectively, these results demonstrated the improvement and robustness of DENA in the detection and quantification of m6A modification in different organisms on transcriptome-wide.
Identification of m6A sites on isoforms of single genes using DENA
Alternative splicing produces isoforms that include or exclude particular exons from the transcripts [3], representing an important mechanism for regulation of gene functions. Previous tools identified and quantified m6A modification by assigning them to genome reference [8, 11, 14]. Thus, they did not distinguish m6A modifications on isoforms produced by alternative RNA splicing. DENA was designed to detect and quantify m6A modification on isoforms from single genes (Fig. 4a). For example, FNR2 (AT1G20020) encodes a leaf-type oxidoreductase in A. thaliana and is present in the chloroplast, having three isoforms based on Araport11 reference. We identified two, three, and two high-confidence m6A sites on the AT1G20020.1, AT1G20020.2, and AT1G20020.3 isoforms, respectively. This result indicated the differential m6A profiles in different isoforms of a single gene and affirmed the capability of DENA in the identification of m6A sites from different isoforms (Fig. 5a).
DENA identified m6A sites on different isoforms of single gene. a Cartoon displays three isoforms of AT1G20020. The different m6A sites on three isoforms are marked with dark bold lines, respectively. The m6A locations on the genome reference are indicated by red inverted triangles. b The distribution of seven high-confidence m6A sites on the three isoforms of AT1G20020 in Col-0, VIRc, and vir-1 lines, respectively. c The changes of m6A rate for the m6A sites in three isoforms of AT1G20020 from Col-0, VIRc, and vir-1 lines, respectively
We next examined them in VIRc and vir-1 lines, and six out of seven m6A sites in three isoforms were shared by both Col-0 and VIRc. However, only three sites were detected in vir-1 (Fig. 5b). For example, for the m6A site that was corresponding to the position 6560974 on the chromosome 1 of TAIR10, the modification rate was 0.59 and 0.42 in isoforms AT1G19000.1 and AT1G19000.2 of Col-0, respectively, which were consistent with the 0.518 and 0.406 in VIRc. In vir-1, however, this site was only identified in AT1G19000.2, with significant reduction of modification rate, only 0.15 (Fig. 5c). Collectively, these results demonstrated the ability of DENA in m6A detection in different isoforms of single genes, which will be helpful in studying the functions of m6A in RNA splicing.
Profiling the m6A modification in mtb and fip37-4 A. thaliana mutants
We used DENA to quantify the m6A modification of three A. thaliana samples, Col-0, mtb, and fip37-4, generated in this study (Additional file 2). We obtained 59,827, 69,504, and 55,134 “RRACH” sites that were supported by at least 50 direct RNA-Seq reads in Col-0, fip37-4, and mtb, respectively. Among them, 19,672 (32.88%), 18,606 (26.77%), and 14,068 (25.52%) m6A sites were identified in Col-0, fip37-4, and mtb, respectively, which showed a high overlap among them (Fig. 6a). The preference for them to be present near the stop codon and within the 3′UTR in transcripts were confirmed (Fig. 6b). We found the modification rates in both fip37-4 and mtb were significantly decreased compared to that in Col-0, agreeing with the LC-MS/MS analysis (Figs. 2a and 6c; Additional file 1: Fig. S4a). In addition, the modification rates were highly correlative between fip37-4 and mtb (Additional file 1: Fig. S4b).
Profiling m6A modification in Col-0, fip37-4, and mtb A. thaliana lines. a Venn diagram shows the common m6A sites among Col-0, fip37-4, and mtb samples generated in this study. b The m6A distribution on transcripts in Col-0, fip37-4, and mtb, respectively. c Jointplot shows the correlation of m6A rate at 11,419 overlapped sites between Col-0 and fip37-4. d Circos shows the profile, sequencing depth, and modification rate of m6A sites in wild-type and m6A-deficient mutant with single-nucleotide resolution, respectively. e The top is the m6A peaks identified in CCA1 using MeRIP-seq in previous study, and the below is the average m6A rate of two high-confidence m6A sites identified by DENA from CCA1 gene in wild-type and m6A-deficient mutant of A. thaliana, respectively. f Point maps the top 25 terms of Gene Ontology enrichment result with all m6A-modified genes. g The changes of average m6A rate at the five high-confidence m6A sites from GIGANTEA gene in wild-type and m6A-deficient mutant of A. thaliana, respectively
To generate a richer resource of m6A modification with single-nucleotide resolution in A. thaliana, we combined all m6A sites identified by DENA in wild-type, VIRc, and three mutants, vir-1, fip37-4, and mtb, and obtained 68,136 m6A sites within 10,223 genes in TAIR10 reference. A clearly reduced average modification rate was observed in the m6A-deficient mutant compared to wild-type (Fig. 6d). For example, two high-confidence m6A sites at position 19248770 and 19248871 were identified by DENA in CCA1 (AT2G46830) of wild-type A. thaliana, which were identified by MeRIP in previous study [18]. However, the average rates were significantly decreased at both sites in the m6A-deficient mutant, especially at the position 19248770 (Fig. 6e). In addition, Liu et al. established a m6A database, named REPIC [28], based on publicly available m6A-seq data. We compared the Arabidopsis m6A sites identified by DENA to the m6A data collected in the REPCI, and 48,744 (71.54%) m6A sites from our model were covered by REPIC.
We performed the Gene Ontology analysis on all m6A-modified genes for functional insights into m6A modification in A. thaliana. These m6A-modified genes were significantly enriched in processes involved in the response to the abiotic and biotic stresses, biosynthetic process, and metabolism in biological process (Fig. 6f). A recent study found that RNA m6A was important for salt stress tolerance in A. thaliana [29]. Consistently, a process of response to salt stress was enriched in our result. Additionally, photosynthesis, light reaction, and electron transport chain were also enriched, in agreement with the previous result that m6A modified transcripts were highly enriched in chloroplast/plastid and protein transport/localization categories [18]. For instance, GIGANTEA (AT1G22770), involved in phytochrome signaling [30] and salt response [29] in A. thaliana, was identified to contain m6A peaks in the 3′UTR [29]. Five high-confidence m6A sites were detected by DENA in its transcripts, including three in the 3′UTR and two in the 9th exon. The modification rates of three sites from 3′UTR and one site from 9th exon were significantly reduced in the m6A-deficient mutant (Fig. 6 g).
Conclusions
m6A is the most abundant modification in mRNA, and it is involved in many aspects of RNA functions. In this study, we developed a neural network, DENA, that can detect and quantify m6A in Nanopore direct RNA-Seq data at single-nucleotide resolution, providing a robust tool for transcriptome-wide profiling of m6A modification.
Using the second-generation sequencing, experimental approaches were developed to detect RNA m6A modification relying on immunoprecipitation [31,32,33,34,35], RNA editing via cytosine deaminase [36], or RNA digestion via m6A-sensitive enzyme [37, 38]. However, they require reverse transcription, specific antibodies or enzymes, and sophisticated experimental procedures, which may introduce biases and variations in their application.
Nanopore direct RNA-Seq can detect electrical signals of RNA m6A modification [32], which overcome these limitations relying on second-generation sequencing-related experimental approaches. One method identified m6A by comparing direct RNA-Seq data of wild-type with that of a matched m6A-deficient or hypomethylated sample [9]. However, the modification rate cannot be accurately quantified, and the m6A-deficient or hypomethylated samples are difficult to generate, representing the major barriers in its application in profiling m6A modification. An alternative approach is to detect m6A modification based on their unique electrical signal fingerprints in direct RNA-Seq data. However, previous attempts were based on the data from in vitro synthetic RNAs in which all “A” residues are replaced by m6A, for training machine learning models [8, 14]. They suffered lower performance on m6A detection in in vivo transcripts as they are likely distorted due to superimposed electrical signals generated by clustered m6A residues in in vitro synthetic sequencing data. By contrast, DENA was developed with the direct RNA-Seq data from in vivo transcripts, and it can detect and quantify m6A at single-nucleotide resolution in the absence of m6A-deficient mutants.
The accuracy and reliability of DENA in m6A detection and quantification were validated in Arabidopsis and human data. On testing direct RNA-Seq data, DENA achieved higher accuracy of m6A identification (90%) in the m6A sites detected by miCLIP in A. thaliana (Fig. 4). Analysis on m6A quantification for known sites in Homo sapiens showed that DENA had a higher correlation with those of the SCARLET method [22] than Nanom6A (Fig. 4). DENA’s better performance may be due to the following reasons: First, DENA was trained on the direct RNA-Seq data containing the naturally occurring m6A sites that are different from the in vitro synthetic data with clustered m6A residues, eliminating the effect of superimposed signals in in vitro data (Fig. 1). Second, a recent study found that the electrical signals of Nanopore sequencing displayed complex heterogeneity in methylation events [39]. Our in vivo training data extracted from over 3000 m6A sites, covered diverse combinations of sequence context, and overcame the limited diversity of the 130 sites containing “RRACH” motif in the in vitro synthetic data. As proposed in previous studies, the performance of m6A detection was improved on the sophisticated neural network, like DENA, by training with our extended in vivo transcribed data.
The inability to identify m6A sites specific to different isoforms of gene transcripts hampered functional studies of m6A in RNA splicing [4]. Another important improvement of DENA is that it can identify and quantify m6A modification on different isoforms of gene transcripts (Fig. 5), which is unavailable in previously methods. DENA assigns sequencing reads to transcript isoforms, which can obtain the isoform compositions and detect the modification sites associated with isoforms. In addition, we evaluated the m6A profiles of fip37-4 and mtb mutants at single-nucleotide resolution using DENA and found a high correlation between the two mutant lines (Additional file 1: Fig. S4b). The results suggested overlapping function between FIP37 and MTB as they are both subunits of the N6-methyltransferase complex catalyzing the formation of m6A modification in RNA.
In summary, DENA (available at https://github.com/weir12/DENA) is the first attempt to use direct RNA-Seq data of in vivo transcribed mRNAs to train neural network model for m6A detection, which overcame the weakness of previous methods trained using in vitro synthetic RNAs with all adenosine residues replaced by m6A. DENA shows a better performance than previous methods and is robust in handing direct RNA-Seq data cross different species. Moreover, we noticed that there were few reads of long noncoding RNAs (lncRNAs) in the human data of direct RNA sequencing. We speculated that most lncRNAs were left out due to lacking poly(A) tail when purifying mRNA by poly(T). Thus, we suggest it may be advantageous to use the poly(A) polymerase to add poly(A) tails to the 3′-end of lncRNAs for detection of m6A modification on lncRNAs, before building the library for direct RNA sequencing. In addition, although our current work was developed using the R9.4 flowcells to perform direct RNA sequencing for m6A detection, our computational framework may be applied to new platforms of direct RNA sequencing technology, which keeps the utility of DENA in pace with future technology development. Collectively, our study provides a useful framework for development of novel models in the detection of other types of RNA modifications using Nanopore direct RNA sequencing.
Methods
Plant materials and growth conditions
All A. thaliana lines were derived from the Columbia (Col-0) accession. Briefly, the hypomorphic allele of FIP37 was generated by a T-DNA insertion within its 7th intron (fip37-4 SALK_018636 allele, termed fip37-4) as reported [17]. The MTB-deficient mutant was generated by rescuing the null mutation through the embryo-specific expression of MTB driven by the ABI3 promoter (ABI3prom:MTB complemented mtb WiscDsLox336H07 allele, termed mtb), because null mutations in MTB are embryonic lethal. Seeds that were stratified at 4 °C for 2 days and were sown on MS medium plates, before they were cultivated in a controlled environment at 22 °C under a 16 h:8 h photoperiod. Seedlings were harvested 14 days after transfer to 22 °C. All materials were frozen immediately after harvest and stored for standby at – 80 °C.
RNA isolation
RNAprep pure Plant Kit (Tiangen Biotech Co., Ltd) was used to extract total RNA from all samples, and Oligotex mRNA Mini Kit was used to purify the polyadenylated mRNA from total RNA. All mRNAs were frozen immediately and were stored at – 80 °C.
LC-MS/MS
LC-MS/MS was performed as described [40]. Briefly, about 500 ng mRNA of each sample was digested by 2 U nuclease P1 (NEB) in 4 μL of 10X Nuclease P1 Reaction Buffer at 37 °C for 30 min, followed by additions of 1 U Thermosensitive Alkaline Phosphatase (1 U/μL) and Thermo Scientific™ FastAP™ reaction buffer. The final mixture incubated at 37 °C for 2 h. The sample was dissolved in 50 μl anhydrous methanol after drying under vacuum, and 10 μL of the solution was used for LC-MS/MS analysis. The samples, and m6A and adenosine (A) standards were separated by reverse-phase ultra-performance liquid chromatography on a C18 column with online mass spectrometry detection using Agilent 6500 QQQ triple-quadrupole LC mass spectrometer in positive electrospray ionization mode. The nucleotides were quantified by using the nucleotide-to-base ion mass transitions of m/z 282.0 to 150.1 (m6A), and m/z 268.0 to 136.0 (A). Three biological replicates for each strain (Col0, mtb, and fip37-4) were performed, and the LC-MS/MS analyses were performed simultaneously on the same machine.
Nanopore direct RNA-Seq and data processing
Nanopore direct RNA sequencing was performed using MinION MkIb with R9.4 flowcells at Nextomics Biosciences Co., Ltd (Wuhan, China). Briefly, mRNA was isolated from about 75 μg total RNA for each sample using the Dynabeads mRNA purification kit (Thermo Fisher Scientific). The quality and quantity of mRNA were assessed using the NanoDrop 2000 spectrophotometer (Thermo Fisher Scientific). Then the nanopore libraries were prepared from 1 μg poly(A) + RNA with the direct RNA sequencing Kit (SQK-RNA001, Oxford Nanopore Technologies) according to the manufacturer’s instructions. The adapter of poly(T) was ligated to the mRNA using T4 DNA ligase (New England Biolabs) in the reaction buffer for 10 min at 25 °C. And then the cDNA was synthesized using SuperScript III Reverse Transcriptase (Thermo Fisher Scientific) with the oligo(dT) adapter and incubate at 50 °C for 50 min, then 70 °C for 10 min, and bring the sample to 4 °C before proceeding to the next step. Then, the RNA-DNA hybrid was purified by Agencourt RNAClean XP magnetic beads (Beckman Coulter). The sequencing adapter was ligated to the mRNA from RNA-DNA hybrid using T4 DNA ligase (New England Biolabs) in the NEBNext Quick Ligation Reaction Buffer (New England Biolabs) for 10 min at 25 °C followed by a second purification step using Agencourt beads (as described above). One microliter reverse-transcribed and adapted RNA were quantified by the Qubit fluorometer DNA HS assay. Then libraries were loaded on GridION using R9.4 flowcells (Oxford Nanopore Technologies) with Library Loading Bead Kit (EXP-LLB001, Oxford Nanopore Technologies) and sequenced using a 48-h run time. Three biological replicates for each strain were sequenced in independent machines and days.
Base-calling was performed with Guppy (version 2.3.4) using default parameters. Reads were aligned to the TAIR10 genome and Araport11 transcriptome references of A. thaliana using minimap2 tool (version 2.813) [41]. Sequence Alignment/Map (SAM) and BAM file manipulations were performed using samtools (version 1.956). The length of direct RNA-Seq reads was calculated using the scripts of wub tool (Oxford Nanopore Technologies Ltd).
Detecting m6A sites in the m6A-dificient mutants using differr
All fast5 files from direct RNA-Seq was bse-called to fastq files by Guppy (version 3.2.4) in this study. The sequence data (fastq files) from m6A-dificient mutants and wild-type A. thaliana were aligned to TAIR10 reference using minimap2. The differential sites between mutant and wild-type using were identified using differr [9] with default parameters. The genomic coordinates were converted to corresponding coordinates of transcriptional isoforms using the R package GenomicFeatures (v1.40.0) [42]. The relative distance is obtained by subtracting the coordinate of the differential sites from that of the core “A” base in the nearest m6A motif “RRACH”. The center “A” residue of “RRACH” that contains differential “error” sites within 5 nucleotides was regarded as m6A modification sites. The electrical signals of direct RNA-Seq data were extracted using Tombo tool [43].
Extracting native training data from direct RNA-Seq reads of A. thaliana
All fast5 files of VIRc and vir-1 samples was base-called to fastq reads by Guppy (version 3.2.4) and subsequently fastq reads were aligned to Araport11 transcriptome reference. We used Tombo re-squiggle algorithm to correct the raw base-calling sequence based on an expected electric level model, and then assigned the corrected base to the raw electric signal segment. After correction, we decided to extract the m6A-modified and unmodified training data from the direct RNA-Seq reads of VIRc and vir-1 A. thaliana lines, respectively. For 3106 common m6A sites identified by differr tool across Cm, Cf, and Vv, we regarded the 11 nucleotides of each site (the nucleotide of m6A site and five nucleotides upstream and downstream) as a “region”, and obtained 3106 “region”. We then extracted the fragments from all base-calling reads at each “region” in VIRc and vir-1, respectively. If a fragment from VIRc sample contained mismatches, we labeled this fragment as a positive event, and extracted its matrix of features (mean, median, standard deviation, dwell time, and base quality) from nanopore electric signals within the “RRACH” window. Conversely, if one fragment from vir-1 mutant did not cause mismatches, we labeled this fragment as negative event, and extracted its matrix of features as above. The dataset containing all matrices of positive events is called m6A-set, while the dataset including all matrices of negative events is called A-set. In all, we obtained the 2251835 matrices of features (mean, median, standard deviation, dwell time, and base quality) from the events of direct RNA-Seq reads, which contains 507,066 in m6A-set and 1,744,769 in A-set.
Training neural network model with the m6A-set and A-set
Bidirectional Long Short-Term Memory (Bi-LSTM) neural network is a variant of Recurrent neural network (RNN) [44]. It can solve the long-term dependency problem (gradient exploding and gradient vanishing) of general RNN and has achieved superior performance in Time Series Prediction, Natural Language Processing, and Machine Translation [44]. Bi-LSTM can capture the rules in the time-series data, such as direct RNA-Seq data, and has been used to detect DNA base modification on Oxford Nanopore sequencing data [15, 16]. Therefore, we consider that Bi-LSTM can be designed to detect of m6A methylation based on nanopore direct RNA sequencing. Then, we divided these labeled features into 12 consensus patterns (“AAACA”, “AAACC”, “AAACT”, “AGACA”, “AGACC”, “AGACT”, “GAACA”, “GAACC”, “GAACT”, “GGACA”, “GGACC”, and “GGACT”) according to the sequence. For each pattern, we further divided its features into training and testing datasets at a ratio of 7:3 for training independent sub-model using Bi-LSTM. In our study, the Bi-LSTM is constructed with three hidden layers after hyperparametric optimization, where bidirectional RNN considers both forward and reverse data flow from neighborhood bases. Each LSTM unit contains multiple hidden nodes. In brief, the flow of tensors during forwarding and backpropagation is visualized in Fig. 3, and the output layer of the Bi-LSTM is flattened and feed into a three-layer fully connected neural network for obtaining the predicted results and calculating the loss function. Samples from different reference sites and reads were randomly shuffled during the training. The prediction labels are scaled by the SoftMax function, and cross-entropy would be minimized to tune the parameters. We use an Adam optimizer with an initial learning rate of 0.0005 to update the weight and bias. However, with the increase of epochs, the learning rate will be reduced. Thus, we decay the learning rate of each parameter group by gamma (0.1) every epoch. To avoid over-fitting, we stop training and retain the model with the best verification set, if three successive epochs do not improve the accuracy in the verification set. The model containing 12 sub-models is the finally available neural network model named DENA. The implementation of DENA relies on the PyTorch, a python-based computing package. The NVIDIA Tesla V100 was used to speed up both the training and testing of our model.
We used the following parameters for evaluating our model:
where TP, FP, TN, FN, and F1 are true positives, false positives, true negatives, false negatives, and F1 score, respectively. We also measure a predictive power of DENA using the area under the curve (AUC) and the receiver operating characteristic curve (ROC) [45]. For DENA model, the input dataset was raw signals from Nanopore direct RNA sequencing. One output result was the predicted modification probability at each “RRACH” site on each nanopore signal, and another output result was the m6A modification rate at each “RRACH” site on reference sequence. The modification rate of each “RRACH” site was calculated as follows:
where Nm and Nu represent the number of signals with a predicted modification probability of not less than 0.5 and the number of signals with a predicted modification probability of less than 0.5 at each site, respectively
Identification and quantification of m6A modification in A. thaliana and human using DENA
After aligning fastq reads to transcriptome reference, the raw signals are assigned to the reference based on an expected electric level model by re-squiggle algorithm of Tombo tool. DENA firstly calculates the position of all “RRACH” motifs in the transcriptome reference, and the following extracts the features from the electric signal. Finally, DENA calculates the depth of sequencing at each position and performs the prediction of both transcriptional coordinate and modification rate (m6A reads/total reads) of m6A from all “RRACH” sites. The sequencing depth of each sample could limit the number of genes and positions that can be identified because of lowly expressed genes being potentially excluded. To maximize the coverage depth of genes and improve the reliability of the m6A ratio, we pool all reads across all replicates of each sample of A. thaliana. For the direct RNA-Seq data of human cells that was established by Ploy N. Pratanwanich et al., we pool all reads across replicate1 (rep1) and replicate3 (rep3) of wild-type, in order to keep the same data size of replicate1 (rep1) of METTL3-knockout mutant cells. To further reduce the impact of sequencing depth and effectively control the false positives, we filtered sites with the following steps: (I) supported by at least 50 reads coverage; (II) the site with modification rate not less than 0.1 as m6A sites; (III) the site with modification rate greater than 0.2 as high-confidence site; (IV) the site with m6A rate between 0.1 and 0.2 as low-confidence site. The coordinate of m6A sites on transcriptome reference was converted to corresponding coordinate on genome reference by in-house python scripts. The consensus motif is plotted by the MEME tool [46], and the distribution of m6A on transcripts is obtained using Guitar package (version 2.8.0) [47]. The m6A profiles of A. thaliana are plotted by Circos software (version 0.69-9). The m6A sites in the profiles are the union of the sites identified by DENA in VIRc, Col-0, and three m6A-deficient mutants of A. thaliana. The average modification rate of m6A site in wild-type A. thaliana was the mean of rates from VIRc and Col-0 samples. The average modification rate of m6A site in m6A-deficient mutant was the mean of rates among three mutants, vir-1, fip37, and mtb. The GRCh38.p13 reference is used for human, and Araport11 and TAIR10 references for A. thaliana. All transcriptome and genome references are downloaded from Ensembl database.
Identification of DENA-predicted m6A modification in A. thaliana using the experimental method
We confirmed the DENA-predicted m6A sites in Arabidopsis using SELECT method described by Xiao et al. [24]. As shown in Fig. 4h, one synthetic DNA probe with PCR adapter (named Dp) and another synthetic DNA probe (named Up) complementarily anneal to RNA but leave a gap of three nucleotides opposite to the candidate site to be identified. As described in other work [26], most mammalian m6A sites are within the G(A/m6A)CU motif. To minimize the over extension of the Up probe by Bst 2.0 DNA polymerase, only dATP, dTTP, dGTP, but not dCTP are added to prevent extension across guanosine in G(A/m6A)CH (H = A, C or U) [26]. If a m6A modification is present in this candidate site of RNA template, it will selectively hinder Bst 2.0 DNA polymerase-mediated single-base elongation of the Up probe and stop it at the “C” base behind m6A-modified base. However, if non-modification presents in the candidate site, the elongation will continue to the candidate “A” base behind the “G” base. Note although this step is not 100% efficient (a small number of elongation products will still reach the m6A-modified site), SplintR ligase will block their connection with Down probes due to the existence of m6A modification at the candidate site in the next ligation step. Meanwhile, we select an unmodified “A” base nearby candidate m6A-modified site as the control. By comparing the cycles of qPCR between the candidate site and the control unmodified “A” base, we can confirm whether there is m6A modification at the candidate site.
In detail, the 5′ phosphorylation of down DNA probe for subsequent ligation was produced by T4 Polynucleotide Kinase within a mixture containing 15 U T4 Polynucleotide Kinase (Vazyme Biotech Co. Ltd.), 50 μM up DNA probe, 0.5 mM ATP, and 1× T4 PNK buffer, at 37 °C for 30 min. Annealing of up and down DNA probes to RNA templates was conducted within the 15 μL mixture containing 1.5 μg total RNAs, 5 pmol 5′ phosphorylation of down DNA probe, 5 pmol up DNA probe, and 0.2 μL RNase inhibitor. The mixture was incubated at 90 °C for 1 min, and then the temperature was reduced at an interval of 10 °C, with 1 min of incubation at each interval end point. After the temperature had dropped to 50 °C, the mixture was kept at 4 °C. Subsequently, the elongation reaction mixture consists of above 15 μL annealed mixture and another 50 μL mixture containing 50 nmol dDTPs (dATP, dGTP, and dTTP) [26], 125 nmol ATP, 0.8 U Bst 2.0 DNA polymerase (NEB), and 0.2 μL RNase inhibitor (Thermo Fisher Scientific) with 1× CutSmart buffer (NEB). The 65 μL mixture for elongation was incubated at 45 °C for 30 min. Finally, a 100 μL ligation reaction mixture consisted of above 65 μL elongation mixture and 35 μL another mixture containing 8 U SplintR ligase (NEB), 20 nmol ATP, 25 μL 50% PEG8000 (RNase-free, Beyotime Biotechnology), and 1× CutSmart buffer. The final mixture was incubated at 37 °C for 40 min, and then was kept at 4 °C. The ligation product was diluted for 100 folds, and then 1 μL was added into a 20-μL scale qPCR reaction with Taq Pro Universal SYBR qPCR Master Mix (Vazyme). The qPCR was performed on QuantStudio 3 Real-Time PCR System with the following parameters: 94 °C, 30 s; (95 °C, 5 s; 57 °C, 20 s; 72 °C, 10 s) × 45 cycles; 4 °C, hold. Data was analyzed using Design and Analysis Setup software (version 2.6). All DNA probes used in this study are listed in Additional file 1: Table S4.
Availability of data and materials
All accessions of datasets used in our study are listed in the Additional file 3. The direct RNA-Seq reads of wild-type, vir-1, and VIR-complemented (VIR::GFP-VIR) A. thaliana lines are downloaded from the European Nucleotide Archive (ENA) under accession PRJEB32782 [9]. The direct RNA-Seq reads of wild-type and METTL3 knockout H. sapiens cells are downloaded from the ENA under accession PRJEB40872 [23]. The direct RNA-Seq reads of in vitro synthetic transcripts are downloaded from the ENA under accession PRJNA511582 [8]. All direct RNA-Seq reads of wild-type, fip37-4, and mtb A. thaliana lines generated by this study have been submitted to the ENA under accession PRJEB45935 [48], and National Genomics Data Center, China National Center for Bioinformation (CNCB-NGDC) under project accession PRJCA007105 and GSA accession CRA005317 [49]. All in-house python scripts used in this study are publicly available as part of DENA that can be download and used from Github with MIT License (https://github.com/weir12/DENA) [50] and Zenodo (https://zenodo.org/record/5603381) [51].
References
Yang Y, Hsu PJ, Chen YS, Yang YG. Dynamic transcriptomic m(6)A decoration: writers, erasers, readers and functions in RNA metabolism. Cell Res. 2018;28(6):616–24. https://doi.org/10.1038/s41422-018-0040-8.
Shi H, Wei J, He C. Where, when, and how: context-dependent functions of RNA methylation writers, readers, and erasers. Mol Cell. 2019;74(4):640–50. https://doi.org/10.1016/j.molcel.2019.04.025.
Black DL. Mechanisms of alternative pre-messenger RNA splicing. Annu Rev Biochem. 2003;72(1):291–336. https://doi.org/10.1146/annurev.biochem.72.121801.161720.
Louloupi A, Ntini E, Conrad T, Orom UAV. Transient N-6-methyladenosine transcriptome sequencing reveals a regulatory role of m6A in splicing efficiency. Cell Rep. 2018;23(12):3429–37. https://doi.org/10.1016/j.celrep.2018.05.077.
Wang X, Lu Z, Gomez A, Hon GC, Yue Y, Han D, et al. N6-methyladenosine-dependent regulation of messenger RNA stability. Nature. 2014;505(7481):117–20. https://doi.org/10.1038/nature12730.
Roundtree IA, Luo GZ, Zhang Z, Wang X, Zhou T, Cui Y, et al. YTHDC1 mediates nuclear export of N(6)-methyladenosine methylated mRNAs. Elife. 2017;6. https://doi.org/10.7554/eLife.31311.
Garalde DR, Snell EA, Jachimowicz D, Sipos B, Lloyd JH, Bruce M, et al. Highly parallel direct RNA sequencing on an array of nanopores. Nat Methods. 2018;15(3):201–6. https://doi.org/10.1038/nmeth.4577.
Liu H, Begik O, Lucas MC, Ramirez JM, Mason CE, Wiener D, et al. Accurate detection of m(6)A RNA modifications in native RNA sequences. Nat Commun. 2019;10(1):4079. https://doi.org/10.1038/s41467-019-11713-9.
Parker MT, Knop K, Sherwood AV, Schurch NJ, Mackinnon K, Gould PD, et al. Nanopore direct RNA sequencing maps the complexity of Arabidopsis mRNA processing and m(6)A modification. Elife. 2020;9. https://doi.org/10.7554/eLife.49658.
Price AM, Hayer KE, McIntyre ABR, Gokhale NS, Abebe JS, Della Fera AN, et al. Direct RNA sequencing reveals m(6)A modifications on adenovirus RNA are necessary for efficient splicing. Nat Commun. 2020;11(1):6016. https://doi.org/10.1038/s41467-020-19787-6.
Jenjaroenpun P, Wongsurawat T, Wadley TD, Wassenaar TM, Liu J, Dai Q, et al. Decoding the epitranscriptional landscape from native RNA sequences. Nucleic Acids Res. 2021;49(2):e7. https://doi.org/10.1093/nar/gkaa620.
Pratanwanich PN, Yao F, Chen Y, Koh CWQ, Wan YK, Hendra C, et al. Identification of differential RNA modifications from nanopore direct RNA sequencing with xPore. Nat Biotechnol. 2021;39(11):1394–402. https://doi.org/10.1038/s41587-021-00949-w.
Lorenz DA, Sathe S, Einstein JM, Yeo GW. Direct RNA sequencing enables m(6)A detection in endogenous transcript isoforms at base-specific resolution. RNA. 2020;26(1):19–28. https://doi.org/10.1261/rna.072785.119.
Gao Y, Liu X, Wu B, Wang H, Xi F, Kohnen MV, et al. Quantitative profiling of N(6)-methyladenosine at single-base resolution in stem-differentiating xylem of Populus trichocarpa using Nanopore direct RNA sequencing. Genome Biol. 2021;22(1):22. https://doi.org/10.1186/s13059-020-02241-7.
Liu Q, Fang L, Yu G, Wang D, Xiao CL, Wang K. Detection of DNA base modifications by deep recurrent neural network on Oxford Nanopore sequencing data. Nat Commun. 2019;10(1):2449. https://doi.org/10.1038/s41467-019-10168-2.
Ni P, Huang N, Zhang Z, Wang DP, Liang F, Miao Y, et al. DeepSignal: detecting DNA methylation state from Nanopore sequencing reads using deep-learning. Bioinformatics. 2019;35(22):4586–95. https://doi.org/10.1093/bioinformatics/btz276.
Shen L, Liang Z, Gu X, Chen Y, Teo ZW, Hou X, et al. N(6)-Methyladenosine RNA modification regulates shoot stem cell fate in Arabidopsis. Dev Cell. 2016;38(2):186–200. https://doi.org/10.1016/j.devcel.2016.06.008.
Luo GZ, MacQueen A, Zheng G, Duan H, Dore LC, Lu Z, et al. Unique features of the m6A methylome in Arabidopsis thaliana. Nat Commun. 2014;5(1):5630. https://doi.org/10.1038/ncomms6630.
Zhong S, Li H, Bodi Z, Button J, Vespa L, Herzog M, et al. MTA is an Arabidopsis messenger RNA adenosine methylase and interacts with a homolog of a sex-specific splicing factor. Plant Cell. 2008;20(5):1278–88. https://doi.org/10.1105/tpc.108.058883.
Ruzicka K, Zhang M, Campilho A, Bodi Z, Kashif M, Saleh M, et al. Identification of factors required for m(6) A mRNA methylation in Arabidopsis reveals a role for the conserved E3 ubiquitin ligase HAKAI. New Phytol. 2017;215(1):157–72. https://doi.org/10.1111/nph.14586.
Anderson SJ, Kramer MC, Gosai SJ, Yu X, Vandivier LE, Nelson ADL, et al. N(6)-methyladenosine inhibits local ribonucleolytic cleavage to stabilize mRNAs in Arabidopsis. Cell Rep. 2018;25(5):1146–57 e1143. https://doi.org/10.1016/j.celrep.2018.10.020.
Liu N, Parisien M, Dai Q, Zheng G, He C, Pan T. Probing N6-methyladenosine RNA modification status at single nucleotide resolution in mRNA and long noncoding RNA. RNA. 2013;19:1848–56.
Pratanwanich PN, Yao F, Chen Y, Koh CWQ, Hendra C, Poon P, et al. Detection of differential RNA modifications from direct RNA sequencing of human cell lines. bioRxiv. 2020:2020.06.18.160010. https://doi.org/10.1101/2020.06.18.160010.
Xiao Y, Wang Y, Tang Q, Wei L, Zhang X, Jia G. An elongation- and ligation-based qPCR amplification method for the radiolabeling-free detection of locus-specific N(6)-methyladenosine modification. Angew Chem Int Ed Engl. 2018;57(49):15995–6000. https://doi.org/10.1002/anie.201807942.
Yang L, Duff MO, Graveley BR, Carmichael GG, Chen LL. Genomewide characterization of non-polyadenylated RNAs. Genome Biol. 2011;12(2):R16. https://doi.org/10.1186/gb-2011-12-2-r16.
Wang Y, Zhang Z, Sepich-Poore C, Zhang L, Xiao Y, He C. LEAD-m(6) A-seq for locus-specific detection of N(6)-methyladenosine and quantification of differential methylation. Angew Chem Int Ed Engl. 2021;60(2):873–80. https://doi.org/10.1002/anie.202007266.
Bujnowska M, Zhang J, Dai Q, Heideman EM, Fei J. Deoxyribozyme-based method for absolute quantification of N (6)-methyladenosine fractions at specific sites of RNA. J Biol Chem. 2020;295(20):6992–7000. https://doi.org/10.1074/jbc.RA120.013359.
Liu S, Zhu A, He C, Chen M. REPIC: a database for exploring the N(6)-methyladenosine methylome. Genome Biol. 2020;21(1):100. https://doi.org/10.1186/s13059-020-02012-4.
Hu J, Cai J, Park SJ, Lee K, Li Y, Chen Y, et al. N(6) -Methyladenosine mRNA methylation is important for salt stress tolerance in Arabidopsis. Plant J. 2021;106(6):1759–75. https://doi.org/10.1111/tpj.15270.
Huq E, Tepperman JM, Quail PH. GIGANTEA is a nuclear protein involved in phytochrome signaling in Arabidopsis. Proc Natl Acad Sci U S A. 2000;97(17):9789–94. https://doi.org/10.1073/pnas.170283997.
Meyer KD, Saletore Y, Zumbo P, Elemento O, Mason CE, Jaffrey SR. Comprehensive analysis of mRNA methylation reveals enrichment in 3’ UTRs and near stop codons. Cell. 2012;149(7):1635–46. https://doi.org/10.1016/j.cell.2012.05.003.
Chen K, Lu Z, Wang X, Fu Y, Luo GZ, Liu N, et al. High-resolution N(6)-methyladenosine (m(6) A) map using photo-crosslinking-assisted m(6) A sequencing. Angew Chem Int Ed Engl. 2015;54(5):1587–90. https://doi.org/10.1002/anie.201410647.
Linder B, Grozhik AV, Olarerin-George AO, Meydan C, Mason CE, Jaffrey SR. Single-nucleotide-resolution mapping of m6A and m6Am throughout the transcriptome. Nat Methods. 2015;12(8):767–72. https://doi.org/10.1038/nmeth.3453.
Koh CWQ, Goh YT, Goh WSS. Atlas of quantitative single-base-resolution N(6)-methyl-adenine methylomes. Nat Commun. 2019;10(1):5636. https://doi.org/10.1038/s41467-019-13561-z.
Shu X, Cao J, Cheng M, Xiang S, Gao M, Li T, et al. A metabolic labeling method detects m(6)A transcriptome-wide at single base resolution. Nat Chem Biol. 2020;16(8):887–95. https://doi.org/10.1038/s41589-020-0526-9.
Meyer KD. DART-seq: an antibody-free method for global m(6)A detection. Nat Methods. 2019;16(12):1275–80. https://doi.org/10.1038/s41592-019-0570-0.
Garcia-Campos MA, Edelheit S, Toth U, Safra M, Shachar R, Viukov S, et al. Deciphering the “m(6)A code” via antibody-independent quantitative profiling. Cell. 2019;178(3):731–47 e716. https://doi.org/10.1016/j.cell.2019.06.013.
Zhang Z, Chen LQ, Zhao YL, Yang CG, Roundtree IA, Zhang Z, et al. Single-base mapping of m(6)A by an antibody-independent method. Sci Adv. 2019;5:eaax0250.
Tourancheau A, Mead EA, Zhang XS, Fang G. Discovering multiple types of DNA methylation from bacteria and microbiome using nanopore sequencing. Nat Methods. 2021;18(5):491–8. https://doi.org/10.1038/s41592-021-01109-3.
Duan HC, Wei LH, Zhang C, Wang Y, Chen L, Lu Z, et al. ALKBH10B is an RNA N(6)-methyladenosine demethylase affecting Arabidopsis floral transition. Plant Cell. 2017;29(12):2995–3011. https://doi.org/10.1105/tpc.16.00912.
Li H. Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics. 2018;34(18):3094–100. https://doi.org/10.1093/bioinformatics/bty191.
Lawrence M, Huber W, Pages H, Aboyoun P, Carlson M, Gentleman R, et al. Software for computing and annotating genomic ranges. PLoS Comput Biol. 2013;9(8):e1003118. https://doi.org/10.1371/journal.pcbi.1003118.
Stoiber M, Quick J, Egan R, Eun Lee J, Celniker S, Neely RK, et al. De novo identification of DNA modifications enabled by genome-guided nanopore signal processing. bioRxiv. 2017:094672. https://doi.org/10.1101/094672;.
Graves A, Schmidhuber J. Framewise phoneme classification with bidirectional LSTM and other neural network architectures. Neural Netw. 2005;18(5-6):602–10. https://doi.org/10.1016/j.neunet.2005.06.042.
Chen X, Wang ZX, Pan XM. HIV-1 tropism prediction by the XGboost and HMM methods. Sci Rep. 2019;9(1):9997. https://doi.org/10.1038/s41598-019-46420-4.
Bailey TL, Boden M, Buske FA, Frith M, Grant CE, Clementi L, et al. MEME SUITE: tools for motif discovery and searching. Nucleic Acids Res. 2009;37(Web Server):W202–8. https://doi.org/10.1093/nar/gkp335.
Cui X, Wei Z, Zhang L, Liu H, Sun L, Zhang SW, et al. Guitar: an R/Bioconductor package for gene annotation guided transcriptomic analysis of RNA-related genomic features. Biomed Res Int. 2016;2016:8367534–8. https://doi.org/10.1155/2016/8367534.
Hang Qin, Liang Ou, Jian Gao, Longxian Chen, Jiawei Wang, Pei Hao, Xuan Li: DENA: training an authentic neural network model using Nanopore sequencing data of Arabidopsis transcripts for detection and quantification of N6-methyladenosine on RNA. ENA BioProject PRJCA007105. hhttps://www.ebi.ac.uk/ena/browser/text-search?query=PRJEB45935. Accessed 9 Dec 2021.
Qin H, Liang O, Gao J, Chen L, Wang J, Hao P, et al. DENA: training an authentic neural network model using Nanopore sequencing data of Arabidopsis transcripts for detection and quantification of N6-methyladenosine on RNA. CNCB-NGDC BioProject. https://ngdc.cncb.ac.cn/gsa/browse/CRA005317;. 2021.
Qin H, Liang O, Gao J, Chen L, Wang J, Hao P, et al. DENA: training an authentic neural network model using Nanopore sequencing data of Arabidopsis transcripts for detection and quantification of N6-methyladenosine on RNA. Github. https://github.com/weir12/DENA. Accessed 16 Aug 2021.
Qin H, Liang O, Gao J, Chen L, Wang J, Hao P, et al. DENA: training an authentic neural network model using Nanopore sequencing data of Arabidopsis transcripts for detection and quantification of N6-methyladenosine on RNA. zenodo. https://zenodo.org/record/5603381. Accessed 27 Oct 2021.
Acknowledgements
We thank the Nextomics Biosciences Co., Ltd for performing Nanopore direct RNA sequencing, and thank Professor Lianfeng Gu for his help in the use of Nanom6A software.
Review history
The review history is available as Additional file 4.
Peer review information
Wenjing She was the primary editor of this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Funding
This work was supported in part by the National Key Research and Development Program of China [2018YFC0310600, 2019YFA0904601], the Strategic Priority Research Program of Chinese Academy of Sciences (XDA24010400), and the National Natural Science Foundation of China (31771412, 31972881).
Author information
Authors and Affiliations
Contributions
X.L., P.H., and J.W. conceived this project and directed the study. L.C. and J.G. assisted H.Q. to perform the experiments. L.O. extracted training data sets and trained the model. H.Q. analyzed data and prepared the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Additional file 1: Fig. S1.
Nanopore direct RNA-Seq implementation and m6A detection with differr tool. Fig. S2. Training DENA. Fig. S3. Confirming the reliability of DENA in m6A quantification. Fig. S4. The correlation of modification rate between wild-type and m6A-deficient A.thaliana mutant. Table S1 Sequencing statistics of poly(A) selected RNAs in biological triplicates from Col0, mtb, and fip37-4 using direct RNA-Seq, respectively. Table S2 The performance of the DENA prediction model that was evaluated with metrics including accuracy, recall, precision, F1-score. Table S3 The comparison of m6A modification rates between DENA and other methods (containing xPore, Nanom6A, SCARLET, LEAD-m6A- seq and Deoxyribozyme-based Method) at the previously identified m6A sites in human. NT: Not detected; -: Not identified; Y: identified as m6A site. Table S4 DNA probes used in the SELECT assay.
Additional file 2.
DENA output containing m6A sites in Col-0, fip37-4 and mtb, and all nonredundant m6A sites identified across all Arabidopsis lines used in this study.
Additional file 3.
List of datasets used in this study.
Additional file 4.
Review history.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Qin, H., Ou, L., Gao, J. et al. DENA: training an authentic neural network model using Nanopore sequencing data of Arabidopsis transcripts for detection and quantification of N6-methyladenosine on RNA. Genome Biol 23, 25 (2022). https://doi.org/10.1186/s13059-021-02598-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s13059-021-02598-3