Figure 1
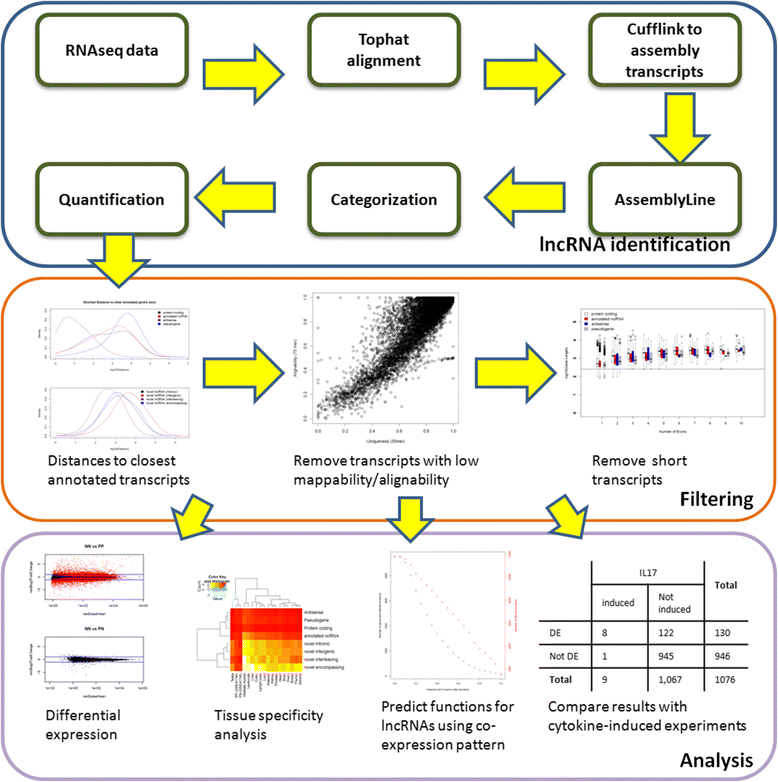
Overview of the analysis pipeline. We first performed Tophat alignment and identified uniquely mapped reads for each RNA-seq sample, we then assembled the transcripts using Cufflinks for each sample. We used a computational approach to nominate potential novel transcripts (Prensner JR et al., [13]) by comparing with Ensembl gene set. We removed those potential novel transcripts which are close (that is, <2 kb) to any exons from any annotated transcripts, inhabited in regions with lower mappability/alignability, or less than 200 bp in length. We quantified the gene expressions using read counts. We then normalized the values across the samples and performed differential expression analysis using DESeq. We inferred the properties and biological functions of the lncRNAs by comparing results with other RNA-seq experiments and using co-expression analysis.
